Welcome to the website of the Iqbal Lab at the Milner Centre for Evolution, Bath!
Welcome to the website of the Iqbal Lab at the Milner Centre for Evolution, Bath!We are a computational research group working on two fronts: half of us study the evolution of bacteria and antibiotic resistance, and half develop new algorithms and computational data structures for comparing and searching through genomes. We love data, algorithms and bacteria! We all come to this from different backgrounds and meet in the middle - we have had microbiologists, mathematicians, computer scientists and physicists in the team, and benefitted from our diversity. We are also proponents of open data, and work collaboratively to build high quality datasets for the benefit of the community.
What's New
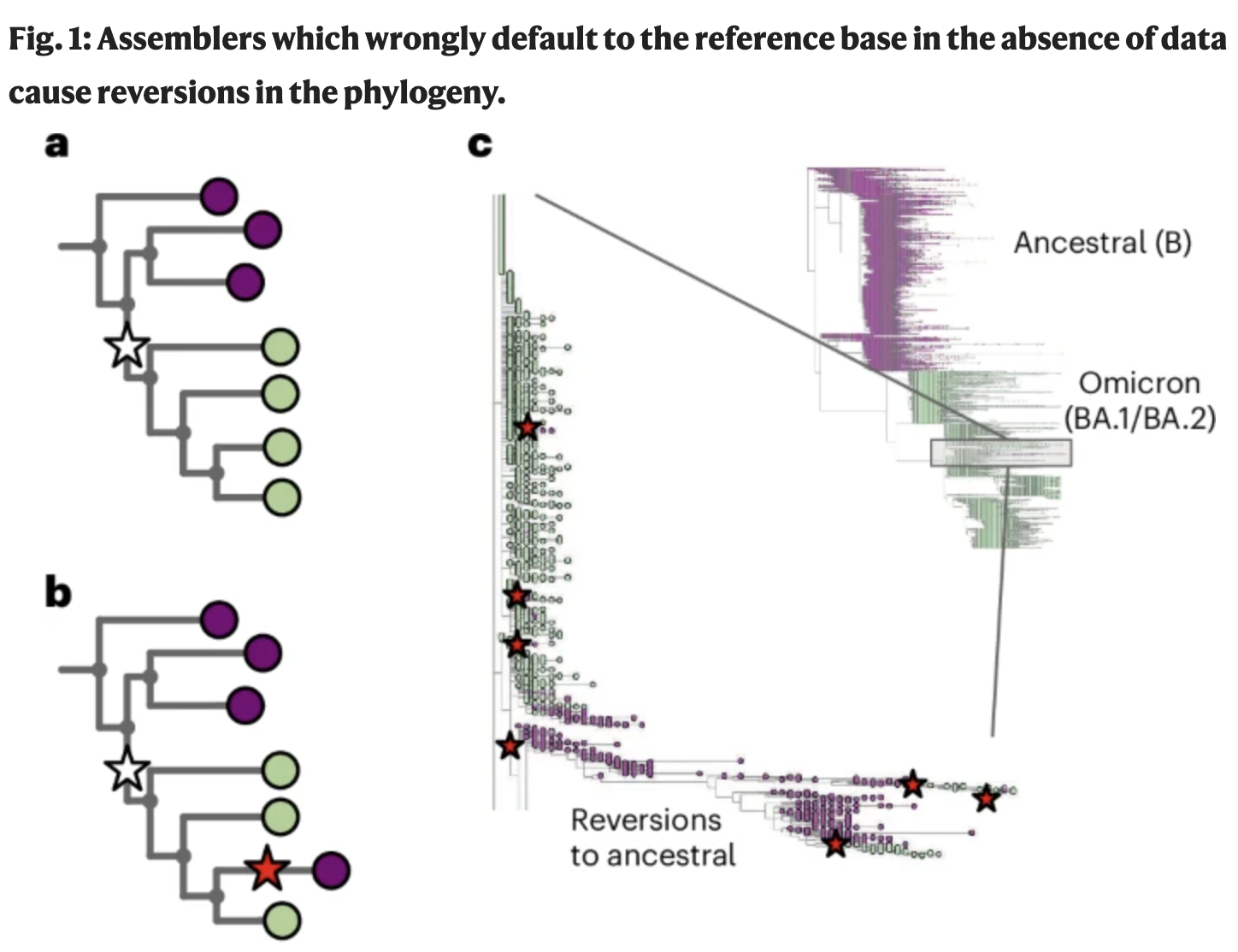
Feb 2026: Our paper on fixing systematic errors in the SARS-CoV-2 phylogeny is published!
Martin Hunt developed a rigorous tiled-amplicon assembler, Viridian, reassembled all open SARS-CoV-2 data and with Russ Corbett-Detig's team constructed a new, better phylogeny. Great collaboration releasing many genomes from the Global South.
Nov 2025: The Genetics Society announces that Zam will be awarded the 2026 Mary Lyon Medal
While this is very welcome, Zam wants to emphasise that he has been lucky to have supportive mentors, and incredibly talented PhD students and postdocs, without whom this would not have happened.
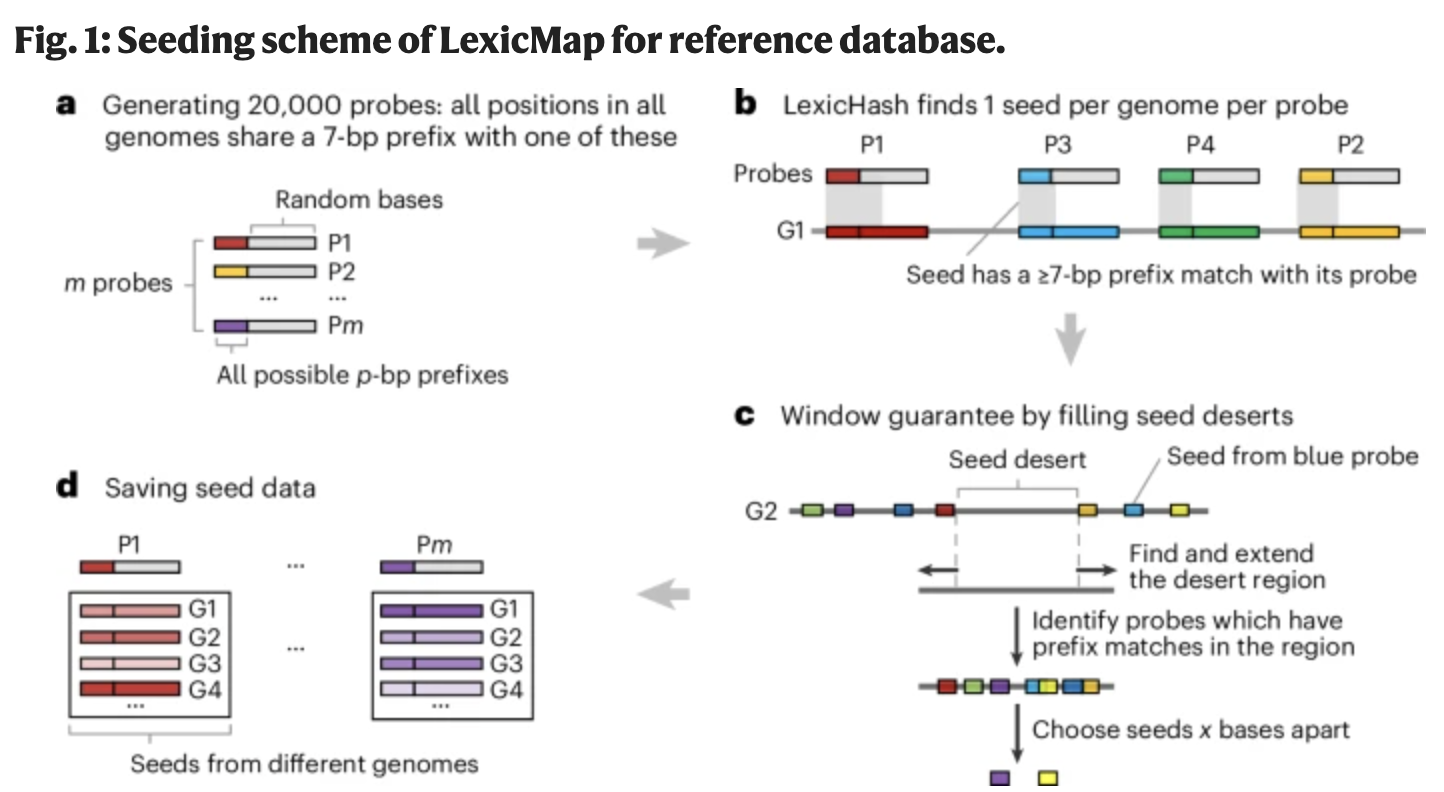
Sept 2025: Our paper on alignment to millions of prokaryotic genomes is published!
New algorithm allows alignment of a gene to 2 million genomes in AllTheBacteria in seconds. Amazing work led by Wei Shen.
Sept 2025: Zam interviewed on the BBC Science in Action radio show
Discussing the Murray Collection study of plasmid evolution; interview starts about 12 minutes in
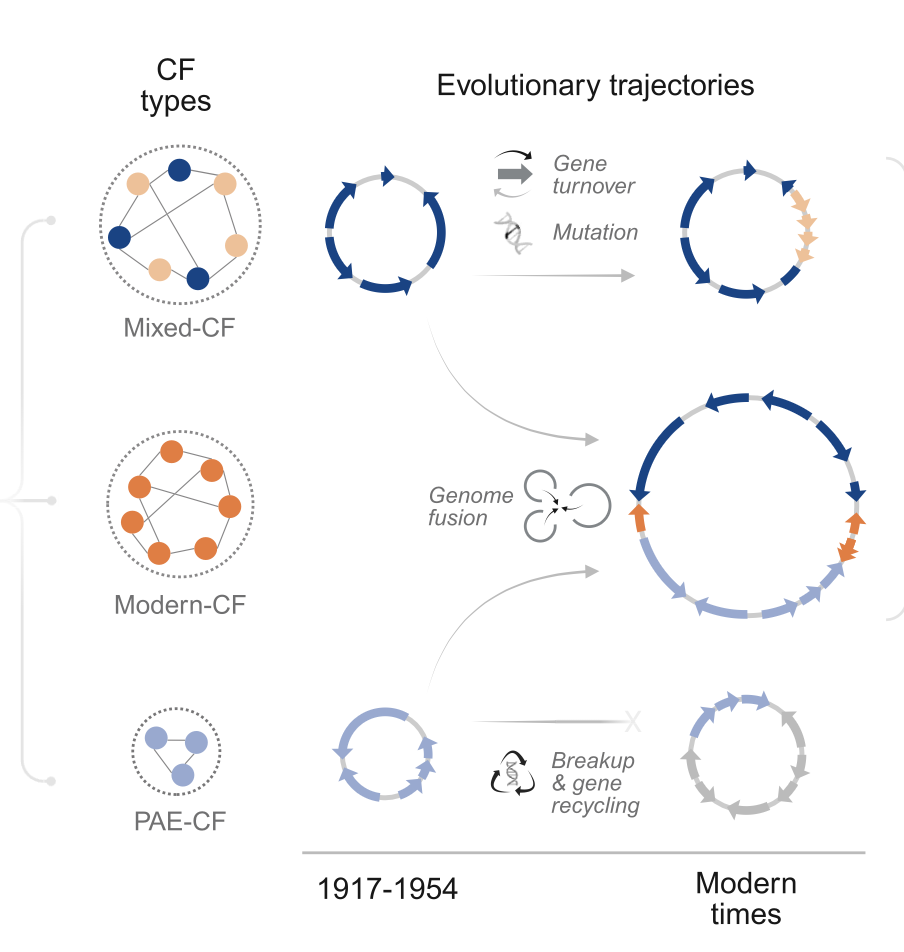
Sept 2025: Our paper on plasmid evolution over the antibiotic epoch is published!
Led by postdoc in the lab Adrian Cazares, collaborative work with Nick Thomson at the Sanger Institute.
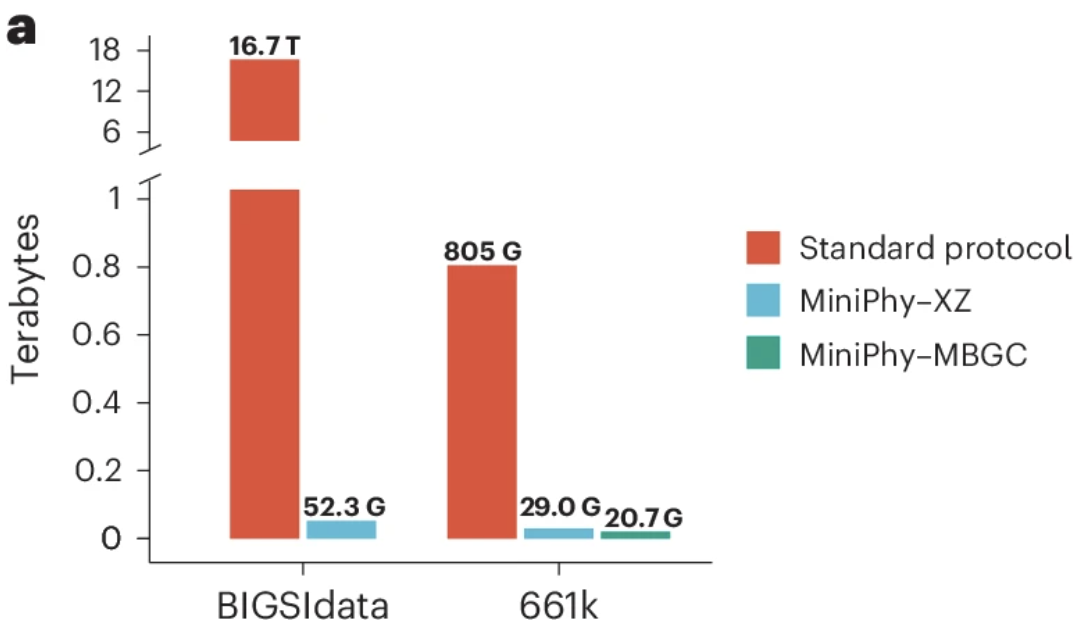
April 2025: Our paper on Phylogenetic Compression is published!
Genomes are not random strings; using evolutionary information to improve compression. Karel Brinda led this, a collaboration also with Mike Baym.